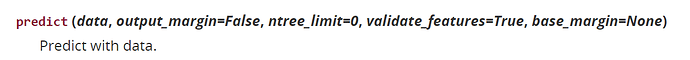Hi all,
I’m unsure if this is the correct place to ask this question,so apologies in advance. I’m using the python implementation of XGboost Pairwise ranking.
The results of my prediction is a list of probabilities, however, I am wondering what the best way is to evaluate such an outcome, or if I made the correct predictions.
I’ve searched multiple online forums but I can’t seem to find a good answer online of how to evaluate the predictions of XGboost Learning to rank.
To illustrate, I’ve followed the python file on github as an example, and it shows:
pred = model.predict(x_test)
When I run my code the outcome is a list of values between 0 and 1. So how do I evaluate how good my predictions are? Is there a built-in way to see what the rankings the predictions have, as compared to the actual rankings?
Again, sorry if this is not the appropriate place to ask such a question.
Thanks in advance.