I am very interested in the split details in xgboost, so I want to plot it.
I can easily plot the decision tree usingsklearn.tree.export_graphviz,
CODE:
#jupyter notebook
from sklearn.tree import export_graphviz
from sklearn.tree import DecisionTreeRegressor
import pydotplus
from IPython .display import Image
def print_graph(clf, feature_names):
"""Print decision tree."""
graph = export_graphviz( clf,
label= "root" ,
proportion= True ,
impurity= False ,
out_file= None ,
feature_names=feature_names,
#class_names={ 0 : "D" , 1 : "R" },
filled= True ,
rounded= True )
graph = pydotplus.graph_from_dot_data(graph)
return Image (graph.create_png())
from sklearn.datasets import load_boston
d = load_boston()
dtr = DecisionTreeRegressor()
dtr.fit(pd.DataFrame(d['data'][:10,:10],columns=d['feature_names'][:10]),d['target'][:10])
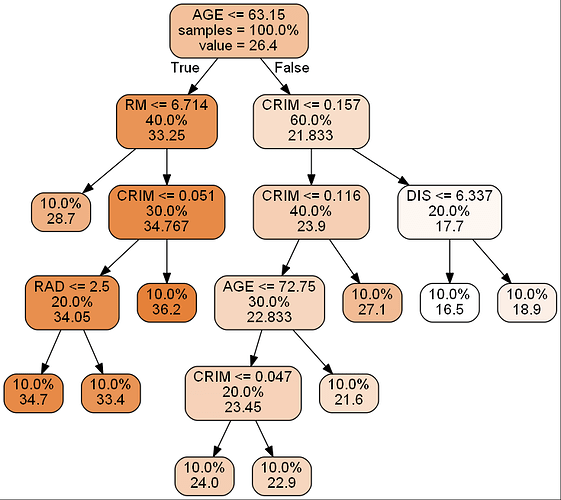
print_graph(dtr,d['feature_names'][:10])
as you can see, in the plot, every split node has 3 values: split condition, sample rate, regresion value.
When I use
xgb.to_graphviz(clf) to plot the tree in xgboost model, I get this picture.#xgbTree|657x500#
I can’t find the sample data amount in the split, after reading the xgb.to_graphviz source code, I know it use
booster.get_dump to get split condition, however , this function still cannot get the explicit information in every split.as the issue Add parameters to the plotting function to control the node shape
Can you help me? how can I plot the xgboost tree as in sklearn ?
Thanks a lot!